The Challenge of Clinical Outcome Prediction
In the high-stakes environment of a hospital, every decision counts. Outcome prediction from clinical text, specifically analyzing admission notes to foresee potential risks, is a critical safety net that can prevent doctors from overlooking rare conditions and help hospitals plan capacities effectively.
The DATEXIS research group has long focused on this challenge, specifically the task of predicting discharge diagnoses based solely on the information available at admission. This creates a „simulated patient“ scenario where decision support is most valuable. The ideal system acts as a second pair of eyes, inferring outcomes based on symptoms, pre-conditions, and risk factors buried in unstructured text.
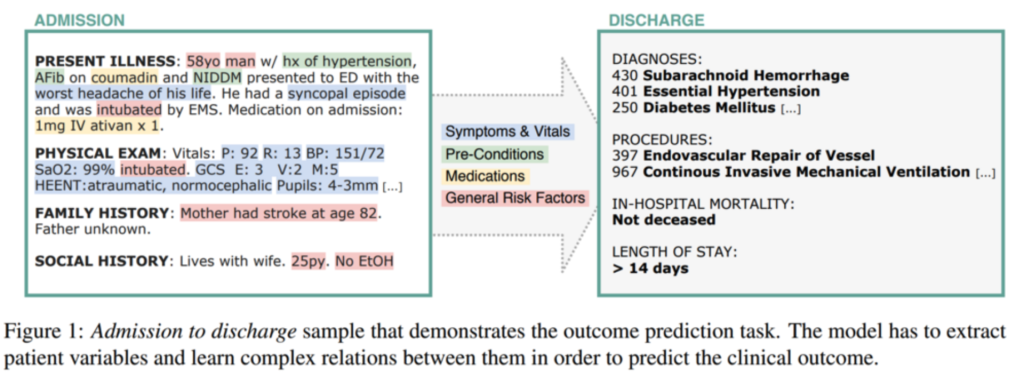

As part of the Appl-FM project („Application-oriented Foundation Models“), our team is now investigating how modern Foundation Models can address these challenges to support demographic change and healthcare efficiency. But are the newest generative models actually better than the specialized tools we already have?
The Era of Encoders: Efficiency and Interpretability
The state-of-the-art in this field is dominated by encoder-based models. A prime example of this efficiency is our CORe model. CORe utilizes a specialized pre-training step to generate Clinical Outcome Representations. By integrating knowledge from public sources like Wikipedia and PubMed, alongside unlabelled clinical notes, the model learns the vital relations between symptoms, risk factors, and outcomes. You can test the model yourself in our demo: https://outcome-prediction.demo.datexis.com/
While models like CORe offer high performance, they historically suffered from a „black box“ problem: they give an answer, but not a reason.
To bridge this trust gap, we developed ProtoPatient, a model that brings interpretability to the forefront. Unlike standard deep learning models, ProtoPatient mimics a doctor’s reasoning process: „This patient looks like that patient.“ It learns prototypical characteristics of diagnoses and bases its predictions on similarities to previous cases, highlighting the specific parts of the text that led to the decision.
Building on this, we introduced SProto, which tackled the „long-tail“ problem, the fact that most diseases are rare and under-represented in data. By using sparse prototypical layers, SProto achieved state-of-the-art performance, proving that specialized encoders could be both accurate and explainable. https://s-proto.demo.datexis.com/
CliniBench: Benchmarking the Generative Shift
With the rapid emergence of Large Language Models (LLMs), the field is shifting. Generative models promise key benefits that encoders lack: zero-shot inference, dynamic label spaces, and the ability to learn from context without expensive retraining. However, their effectiveness in real-world clinical settings has been largely untested.
To answer this, we developed CliniBench, the first benchmark designed to systematically compare generative LLMs against encoder-based classifiers for diagnosis prediction. This work has been accepted at EACL 2026, where we will present our findings.
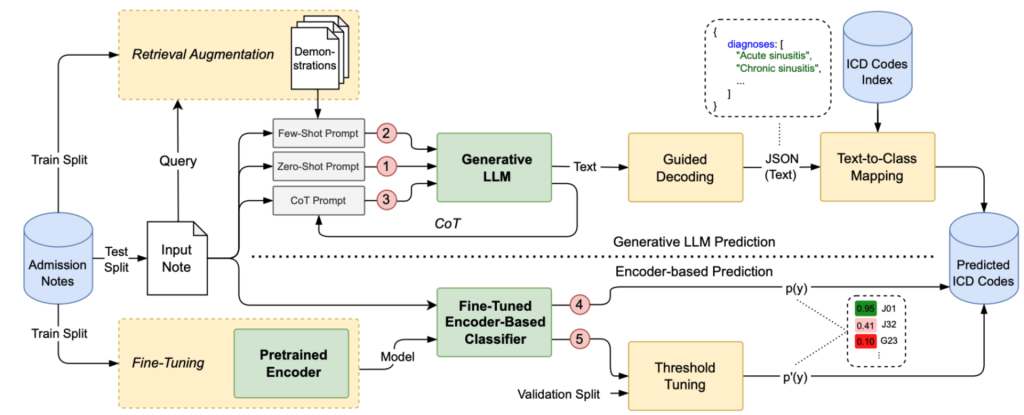
The results were surprising. Despite the hype, encoder-based classifiers consistently outperformed generative models in raw diagnosis prediction. While LLMs are flexible, they are prone to hallucinations and high inference costs. However, we found that „In-Context Learning“ strategies, such as Retrieval-Augmented Generation (RAG), significantly boosted LLM performance. This suggests that while GenAI isn’t ready to replace encoders yet, it holds immense potential for resource-constrained environments where training data is scarce.
The Future: Modular Agentic Models That „Think“ and Verify
For the future we envision not a single monolithic system but a set of modular agents. This modular approach uses a central AI orchestrator to coordinate multiple specialized language models, each contributing distinct strengths (reasoning, questioning, verification) to the diagnostic process. Instead of relying on a single monolithic model, the system dynamically assigns subtasks, like deciding what information to gather next or how to evaluate test results, to the most suitable model.
This design improves accuracy, transparency, and efficiency while making it easier to upgrade or swap individual components as models improve. We envision this multi-agent diagnostic AI designed to gather patient history and generate structured medical summaries, differential diagnoses, and management plans, without giving individualized medical advice on its own.
Crucially, this process relies on medical verifiers. We are exploring how to automate the verifier creation process grounded on knowledge bases, such as WikiDoc. By validating the model’s „thought process“ against established medical knowledge, rather than just checking the final answer, we can ensure that the AI is not only accurate but also plausible and trustworthy.
Conclusion
The transition from specialized encoders to general-purpose Foundation Models is the next frontier in predictive medicine. While benchmarks like CliniBench show that encoders currently hold the crown for pure performance, the adaptability of generative models is undeniable. By equipping these models with the ability to orchestrate complex reasoning tasks and verify their own logic, we are paving the way for AI that does not just predict the future but explains it, building the trust necessary for true clinical adoption.


